This is the first post in a series of blogs exploring blood-based biosurveillance for novel pathogen detection as part of the NAO’s effort to evaluate different biosurveillance approaches. We thank our colleagues at the NAO for their valuable feedback, which has significantly improved this article.
Introduction
Detecting novel viral pathogens remains a critical challenge in public health. To address this challenge, biosurveillance systems must carefully select appropriate sample types for analysis. This choice significantly influences detection cost, sensitivity, and the range of pathogens that can be identified (1–3). The Nucleic Acid Observatory has previously examined a variety of sample types, including wastewater, air, and pooled swabs. This blog series investigates blood as a potential sample type for large-scale disease monitoring.
Blood, formally referred to as whole blood, and its components, such as plasma, represent promising targets for biosurveillance. They contain diverse biomarkers of infection, including antibodies, intact pathogens, pathogen-derived proteins, and nucleic acids (4), which can serve as the targets for molecular assays. Moreover, well-established infrastructure exists for blood collection and testing (5,6). Several existing systems could potentially be leveraged for early detection of emerging pathogens, offering complementary approaches to other surveillance methods. These include: the blood supply system (developed primarily for transfusion medicine and plasma-derived therapies) (7); blood biobanks; and samples collected for diagnostic testing.
In this post, we explore the composition and characteristics of blood as well as the presence of viruses within it. We introduce two distinct sample types, namely whole blood and plasma. Future posts will examine promising sampling strategies in the context of existing blood and plasma collection systems, as well as the sensitivity of blood and plasma sampling for pathogen detection.
Composition and characteristics of blood
Blood consists of two components: formed elements and plasma (Figure 1). Formed elements are cells or cell fragments that originate from hematopoietic stem cells within the bone marrow. They encompass red blood cells, white blood cells, and platelets, together constituting approximately 37-54% of blood volume (8,9). Plasma, constituting the remaining fraction, is the fluid medium that transports formed elements, along with nutrients, waste products, proteins, and microbes (8,9). Serum refers to plasma from which proteins involved in clotting have been removed (10).

Blood serves as the body’s primary circulatory medium, transporting oxygen, nutrients, and metabolic waste throughout the body (9). It also serves as a dynamic repository of health information. In clinical settings, blood analysis is crucial for diagnosing and monitoring a wide range of conditions, including infections, cancer (11), cardiovascular diseases (12), metabolic disorders (13), and genetic abnormalities (14). It carries indicators of infection and immune activity, including intact pathogens, pathogen-derived proteins and nucleic acids, leukocytes, and antibodies (15). Some of these biomarkers can be detected in blood before the onset of symptoms or persist long after the acute phase of an infection (7).
Viruses in blood
Entry into the bloodstream
While many viruses primarily infect specific tissues, some can enter the bloodstream, either as their main target or during the course of infection. When viruses do enter the blood, this can occur through several mechanisms:
- Direct introduction through breaks in physical barriers. This includes entry via needle sticks, insect bites, or damaged skin, as well as through mucous membranes in the respiratory, gastrointestinal, and genital tracts.
- Spread from local infections. As infections progress, viruses can overwhelm local immune responses and enter the bloodstream. This can occur through damaged blood vessels, infection of endothelial cells lining the vessels, or via the lymphatic system.
- Sexual transmission. This route involves specific adaptations for transmission via genital mucosa and fluids.
- Vertical transmission from mother to fetus through the placenta.
The lymphatic system can serve as a gateway for viruses to enter the bloodstream (16). Many viral infections begin in epithelial cells of exposed tissues, such as the skin or respiratory tract. From these primary infection sites, viral particles or infected cells can enter nearby lymphatic vessels and migrate to lymph nodes. This lymphatic transport is often part of a healthy immune response, where dendritic cells capture viral antigens and migrate to lymph nodes to present them to immune cells.
Lymph nodes, rich in immune cells like lymphocytes and monocytes, are connected to the circulatory system. While the immune response often contains infections within lymph nodes, some viruses can breach this defense. Certain viruses, like HIV and measles, can infect and replicate within immune cells in the lymph nodes, potentially amplifying the infection (17,18). In cases where viruses overcome lymph node defenses, they may enter the bloodstream either as free-floating particles or within infected immune cells, enabling systemic spread throughout the body (19).
Viremia patterns and viral persistence
Once viruses enter the bloodstream, they can spread throughout the body in a process known as hematogenous spread. The presence of infectious virus particles in the blood is termed viremia, which can be classified as either active or passive. Active viremia results from viral reproduction within the host, while passive viremia occurs when virus particles are introduced into the blood without local replication, such as through the bite of an infected mosquito (19).
Viremia typically progresses through two phases: primary and secondary (19,20). Primary viremia refers to the initial release of virus particles into the blood following reproduction at the entry site. At this stage, viral concentrations are usually low. Secondary viremia occurs when the virus disseminates to other tissues, replicates, and is subsequently released back into the bloodstream in higher concentrations. This secondary phase often elicits more severe immune responses and symptoms.
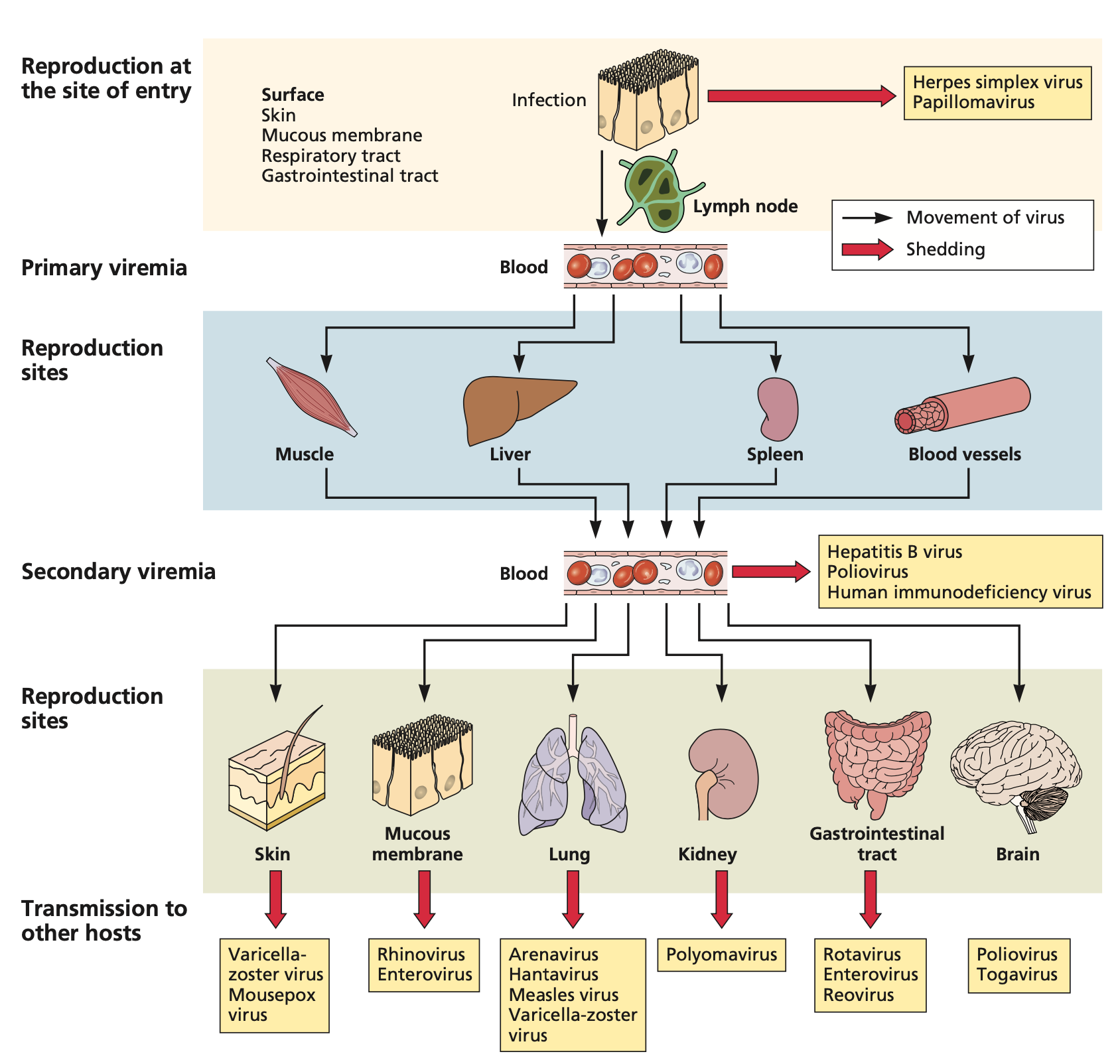 Figure 2: Entry, dissemination, and shedding of viruses in the bloodstream. Adapted from Flint et al. (19), Principles of Virology, 5th Ed., 2020. Reproduced with permission from John Wiley & Sons via Copyright Clearance Center.
Figure 2: Entry, dissemination, and shedding of viruses in the bloodstream. Adapted from Flint et al. (19), Principles of Virology, 5th Ed., 2020. Reproduced with permission from John Wiley & Sons via Copyright Clearance Center.
However, viruses exhibit diverse life cycles and host interactions, resulting in varied patterns of bloodstream presence. For some viruses, such as dengue (21) or Zika (22), viremia is a normal part of the infection cycle and is typically present in all infected individuals. These arboviruses rely on bloodstream circulation to facilitate transmission via mosquito vectors. Similarly, bloodborne viruses like HIV, hepatitis B, and hepatitis C can establish persistent infections, maintaining continuous low-level viral production in the bloodstream as a key part of their lifecycle and transmission strategy (23). HIV, for instance, actively infects CD4+ T cells and macrophages in the bloodstream, maintaining chronic infection through integration into the host genome (18). Certain viruses, like HIV and Epstein-Barr virus, can also establish latent infections with periods of viral silence punctuated by reactivation and intermittent viremia (18,24).
In contrast, for respiratory viruses like influenza, rhinovirus, or SARS-CoV-2, the detection of viral RNA or infectious viral particles in blood is generally rare and often indicates a more severe or systemic spread of the infection. In cases where viral RNA does enter the blood, it typically becomes undetectable after a period of days or a few weeks (< 4 weeks) (25–28). Studies on SARS-CoV-2 found viral RNA in blood samples from around 10% of infected individuals, with higher rates in severely ill patients (25). Similarly, rhinovirus RNA was detected in blood samples from 15% of severely immunocompromised patients with high viral loads in respiratory samples (26), as well as in about 12% of blood samples from children with rhinovirus-positive nasopharyngeal swabs and severe respiratory infections (29). Influenza A viral RNA is even rarer in blood, with no reliable detection even in symptomatic patients during peak flu season (30). However, for pandemic influenza A/H1N1/2009, approximately 10% of hospitalized patients tested positive for viral RNA in their blood, with its presence strongly associated with more severe symptoms (28). For respiratory pathogens, the association between viral nucleic acid in blood and severe disease means affected individuals are unlikely to be blood or plasma donors outside of diagnostic testing, as donation centers typically screen out symptomatic people.
While these findings suggest that viral nucleic acids from respiratory pathogens can enter the bloodstream, their presence doesn’t necessarily indicate infectious virus. For the purposes of this article, we performed only a shallow literature review; a more comprehensive literature review could further elucidate the dynamics of true viremia versus viral nucleic acids in blood across different pathogens and populations.
Clearance
Multiple mechanisms contribute to viral clearance from the bloodstream. The mononuclear phagocyte system, comprising phagocytic cells in the liver, spleen, and lymph nodes, engulfs and destroys circulating viruses (19). As the immune response develops, antibodies bind to viral particles, forming antibody-virus complexes that are more readily recognized and cleared by these phagocytic cells. The liver and kidneys are key organs in this process: the liver filters blood and can excrete viral components into bile, which is then eliminated through feces, while the kidneys may filter smaller viral particles or fragments into urine. The exact mechanisms and efficiency of viral clearance vary depending on the specific virus and the stage of infection.
Plasma or whole blood?
Whole blood and plasma are distinct sample types with different physical properties relevant to viral nucleic acid detection. This section examines these characteristics, providing a foundation for understanding their potential in novel pathogen detection. While physical properties are important, other factors such as sampling strategies and detection sensitivity also influence their suitability for biosurveillance, which will be explored in future posts.
Viral nucleic acids can be present in blood in three primary forms:
- Within blood cells: Blood-borne pathogens like HIV-1, human parvovirus B19, and Epstein-Barr virus primarily infect and replicate within specific blood cells. In these cells, viral nucleic acids may exist as encapsidated particles, integrated into the host genome (provirus), or as free-floating genetic material in the cell’s cytoplasm or nucleus (31).
- As intact virions in plasma: Complete virus particles can circulate freely in plasma. Some viruses, such as hepatitis B and C, may associate with clotting proteins or adhere to blood cell surfaces while in plasma (32).
- As cell-free nucleic acids in plasma: Viral genetic material can be found free-floating in plasma, even in the absence of intact virions. These cell-free viral nucleic acids originate from lysis of virus-infected cells, degradation of virions, and active secretion by viable cells (4,33,34).
Whole blood captures viral nucleic acids in all these forms, potentially offering more comprehensive coverage. This includes detection of viruses primarily residing within blood cells and those establishing latent infections. However, whole blood samples contain significantly more human DNA than plasma, which can impact the sensitivity of pathogen detection via metagenomic sequencing. Emerging approaches, such as CRISPR-based depletion of human reads, show promise in addressing this challenge (35).
Plasma contains viral nucleic acids in the form of free-floating genetic material and intact virions. While plasma lacks the intracellular viral content found in whole blood, it plays a crucial role in viral dissemination. Viruses that propagate in or via the bloodstream must release particles or nucleic acids into plasma, which should enable their detection. However, many pathogens do not routinely appear in blood, in which case they will be missing from both whole blood and plasma. Ultimately, the presence and detectability of a virus in plasma depends on factors such as its tissue tropism, replication cycle, ability to establish systemic infection, and the timing relative to symptom onset.
Conclusion
Blood-based biosurveillance presents a potentially valuable approach for early detection of novel pathogens, leveraging existing infrastructure and the information-rich nature of blood. In this post, we covered the composition and characteristics of blood, as well the dynamics of viruses in the bloodstream. Future work will explore promising sampling strategies within existing large-scale collection systems for both whole blood and plasma. We also plan to analyze the relative abundance of human-infecting pathogens in blood metagenomic sequencing data and integrate these findings with estimates of viral prevalence or incidence in the community, similar to our previous analyses for wastewater and swab sampling. These investigations will help determine the viability and potential impact of integrating blood-based surveillance into a comprehensive early warning system for emerging pandemic threats.
References
- Gauthier NPG, Chorlton SD, Krajden M, Manges AR. Agnostic Sequencing for Detection of Viral Pathogens. Clin Microbiol Rev. 2023 Mar 23;36(1):e0011922.
- Bradshaw W, Grimm S. Comparing sampling strategies for early detection of stealth biothreats [Internet]. 2023 [cited 2024 Jun 7]. Available from: https://naobservatory.org/reports/comparing-sampling-strategies-for-early-detection-of-stealth-biothreats
- Santarpia JL, Klug E, Ravnholdt A, Kinahan SM. Environmental sampling for disease surveillance: Recent advances and recommendations for best practice. Journal of the Air & Waste Management Association. 2023 Jun 3;73(6):434–61.
- Loy C, Ahmann L, De Vlaminck I, Gu W. Liquid Biopsy Based on Cell-Free DNA and RNA. Annu Rev Biomed Eng [Internet]. 2024 Feb 12; Available from: http://dx.doi.org/10.1146/annurev-bioeng-110222-111259
- Baş S, Carello G, Lanzarone E, Ocak Z, Yalçındağ S. Management of blood donation system: Literature review and research perspectives. In: Springer Proceedings in Mathematics & Statistics. Cham: Springer International Publishing; 2016. p. 121–32. (Springer proceedings in mathematics & statistics).
- Etherington C, Palumbo A, Holloway K, Meyer S, Labrecque M, Rubini K, et al. Barriers and enablers to and strategies for promoting domestic plasma donation throughout the world: Overarching protocol for three systematic reviews. PLoS One. 2023 Dec 21;18(12):e0296104.
- McCullough JJ, editor. Transfusion medicine. Fifth edition. Hoboken, NJ Chichester: Wiley Blackwell; 2021. 580 p.
- Maue-Dickson W, Dickson DR. Anatomy and physiology related to cleft palate: current research and clinical implications. Plast Reconstr Surg. 1980;65(1):83–90.
- Gordon Betts J, Young KA, Wise JA, Johnson E, Poe B, Kruse DH, et al. Anatomy and Physiology 2e. Houston, Texas: OpenStax; 2022. (Open Touro Adopted).
- Krebs HA. Chemical Composition of Blood Plasma and Serum. Annu Rev Biochem. 1950 Jun;19(1):409–30.
- Crowley E, Di Nicolantonio F, Loupakis F, Bardelli A. Liquid biopsy: monitoring cancer-genetics in the blood. Nat Rev Clin Oncol. 2013 Aug 9;10(8):472–84.
- Vasan RS. Biomarkers of cardiovascular disease: Molecular basis and practical considerations. Circulation. 2006 May 16;113(19):2335–62.
- Guerrero RB, Salazar D, Tanpaiboon P. Laboratory diagnostic approaches in metabolic disorders. Ann Transl Med. 2018 Dec;6(24):470.
- Sehnert AJ, Rhees B, Comstock D, de Feo E, Heilek G, Burke J, et al. Optimal detection of fetal chromosomal abnormalities by massively parallel DNA sequencing of cell-free fetal DNA from maternal blood. Clin Chem. 2011 Jul;57(7):1042–9.
- Marshall JS, Warrington R, Watson W, Kim HL. An introduction to immunology and immunopathology. Allergy Asthma Clin Immunol. 2018 Sep 12;14(2):49.
- Nash AA, Dalziel RG, Ross Fitzgerald J. Mims’ pathogenesis of Infectious Disease. San Diego, CA: Academic Press; 2015. 365 p.
- Rota PA, Moss WJ, Takeda M, de Swart RL, Thompson KM, Goodson JL. Measles. Nat Rev Dis Primers [Internet]. 2016 Jul 14;2(16049). Available from: http://dx.doi.org/10.1038/nrdp.2016.49
- Bekker LG, Beyrer C, Mgodi N, Lewin SR, Delany-Moretlwe S, Taiwo B, et al. HIV infection. Nat Rev Dis Primers. 2023 Aug 17;9(1):42.
- Flint J, Racaniello VR, Rall GF, Hatziioannou T, Skalka AM. Principles of virology, Volume II. 5th ed. Washington, D.C., DC: American Society for Microbiology; 2020. 1136 p. (ASM Books).
- Burrell CJ, Howard CR, Murphy FA. Pathogenesis of Virus Infections. Fenner and White’s Medical Virology. 2017;77–104.
- Guzman MG, Gubler DJ, Izquierdo A, Martinez E, Halstead SB. Dengue infection. Nat Rev Dis Primers. 2016 Aug 18;2(1):16055.
- Plourde AR, Bloch EM. A literature review of Zika virus. Emerg Infect Dis. 2016 Jul;22(7):1185–92.
- Boldogh I, Albrecht T, Porter DD. Persistent Viral Infections. University of Texas Medical Branch at Galveston; 1996.
- Hoover K, Higginbotham K. Epstein-Barr Virus. In: StatPearls [Internet]. StatPearls Publishing; 2023.
- Andersson MI, Arancibia-Carcamo CV, Auckland K, Baillie JK, Barnes E, Beneke T, et al. SARS-CoV-2 RNA detected in blood products from patients with COVID-19 is not associated with infectious virus. Wellcome Open Res. 2020 Oct 12;5:181.
- Van Rijn AL, Claas EC, von dem Borne PA, Kroes ACM, de Vries JJC. Rhinovirus viremia in adult patients with high viral load in bronchoalveolar lavages. J Clin Virol. 2017 Nov 1;96:105–9.
- Ng LFP, Wong M, Koh S, Ooi EE, Tang KF, Leong HN, et al. Detection of Severe acute respiratory syndrome Coronavirus in blood of infected patients. J Clin Microbiol. 2004 Jan;42(1):347–50.
- Tse H, To KKW, Wen X, Chen H, Chan KH, Tsoi HW, et al. Clinical and virological factors associated with viremia in pandemic influenza A/H1N1/2009 virus infection. PLoS One. 2011 Sep 27;6(9):e22534.
- Fuji N, Suzuki A, Lupisan S, Sombrero L, Galang H, Kamigaki T, et al. Detection of human rhinovirus C viral genome in blood among children with severe respiratory infections in the Philippines. PLoS One. 2011 Nov 8;6(11):e27247.
- Stramer SL, Collins C, Nugent T, Wang X, Fuschino M, Heitman JW, et al. Sensitive detection assays for influenza RNA do not reveal viremia in US blood donors. J Infect Dis. 2012 Mar 15;205(6):886–94.
- Mims. The Spread of Microbes Through blood. In 2001.
- Tu T, Ajoyan H, Nur Umami R, Veeraraghavan V, Boldbaatar D, Najim MAM, et al. Inhibition of Cellular Factor TM6SF2 Suppresses Secretion Pathways of Hepatitis B, Hepatitis C, and Hepatitis D Viruses. J Infect Dis. 2024 Feb 26;jiae098.
- Bronkhorst AJ, Ungerer V, Oberhofer A, Gabriel S, Polatoglou E, Randeu H, et al. New Perspectives on the Importance of Cell-Free DNA Biology. Diagnostics (Basel) [Internet]. 2022 Sep 3;12(9). Available from: http://dx.doi.org/10.3390/diagnostics12092147
- Szilágyi M, Pös O, Márton É, Buglyó G, Soltész B, Keserű J, et al. Circulating Cell-Free Nucleic Acids: Main Characteristics and Clinical Application. Int J Mol Sci [Internet]. 2020 Sep 17;21(18). Available from: http://dx.doi.org/10.3390/ijms21186827
- Chan AP, Siddique A, Desplat Y, Choi Y, Ranganathan S, Choudhary KS, et al. A CRISPR-enhanced metagenomic NGS test to improve pandemic preparedness. Cell Rep Methods. 2023 May 22;3(5):100463.