Summary: The NAO will increase our sequencing significantly over the next few months, funded by a $3M grant from Open Philanthropy. This will allow us to scale our early-warning system to where we could flag many engineered pathogens early enough to mitigate their worst impacts, and also generate large amounts of data to develop, tune, and evaluate our detection systems.
One of the biological threats the NAO is most concerned with is a ‘stealth’ pathogen, such as a virus with the profile of a faster-spreading HIV. This could cause a devastating pandemic, and early detection would be critical to mitigate the worst impacts. If such a pathogen were to spread, however, we wouldn’t be able to monitor it with traditional approaches because we wouldn’t know what to look for. Instead, we have invested in metagenomic sequencing for pathogen-agnostic detection. This doesn’t require deciding what sequences to look for up front: you sequence the nucleic acids (RNA and DNA) and analyze them computationally for signs of novel pathogens.
We’ve primarily focused on wastewater because it has such broad population coverage: a city in a cup of sewage. On the other hand, wastewater is difficult because the fraction of nucleic acids that come from any given virus is very low,1 and so you need quite deep sequencing to find something. Fortunately, sequencing has continued to come down in price, to under $1k per billion read pairs. This is an impressive reduction, 1/8 of what we estimated two years ago when we first attempted to model the cost-effectiveness of detection, and it makes methods that rely on very deep sequencing practical.
Over the past year, in collaboration with our partners at the University of Missouri (MU) and the University of California, Irvine (UCI), we started to sequence in earnest:
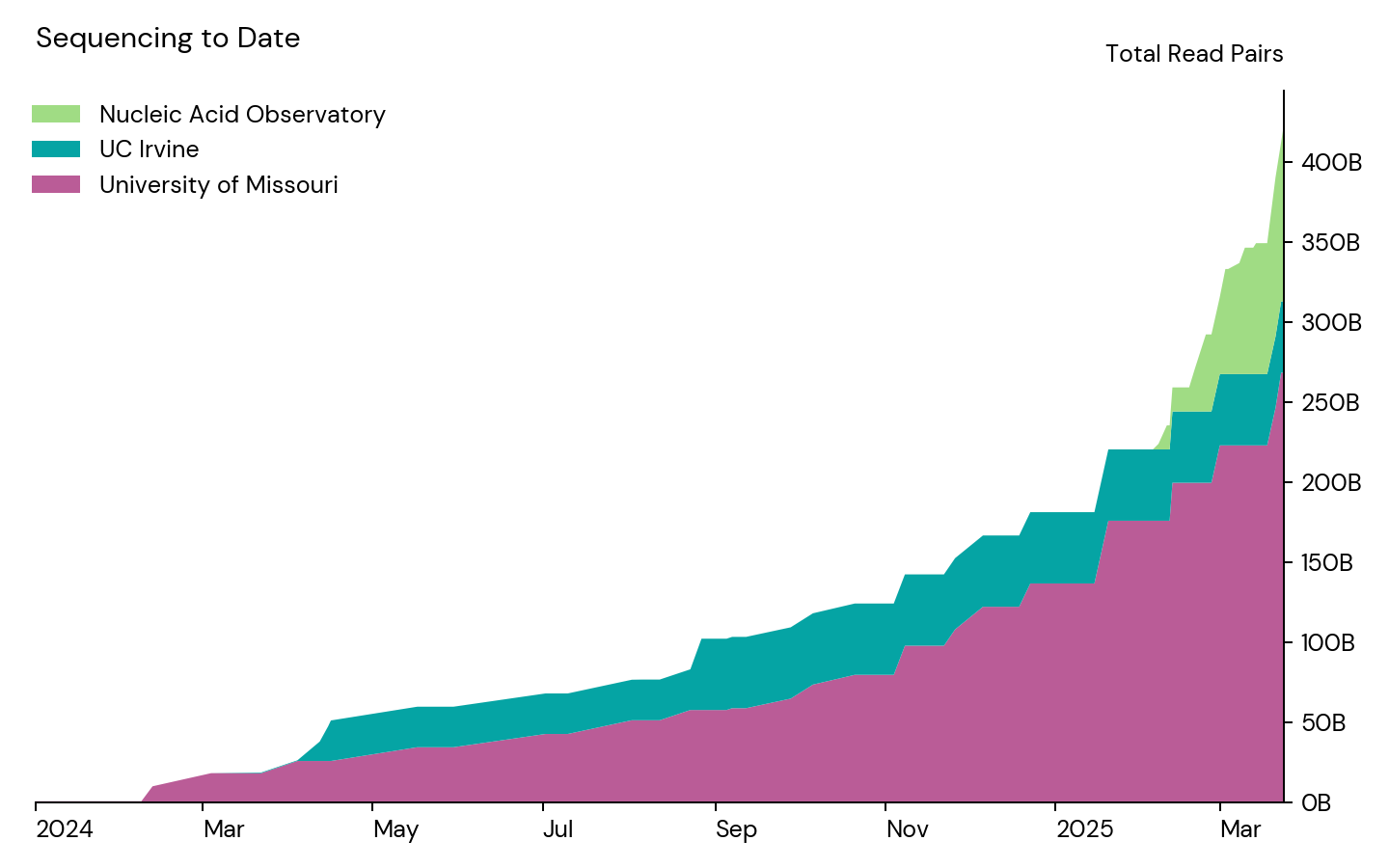
We believe this represents the majority of metagenomic wastewater sequencing produced in the world to date, and it’s an incredibly rich dataset. It has allowed us to develop and test our algorithms for pathogen identification, and we’re eager to share it with others who are working to develop their own computational approaches to this problem. This is a valuable start, and is enough to provide very sensitive coverage of gastrointestinal viruses. To get a pilot early warning system to where it could usefully flag other viruses, however, we’ll need to ramp up sequencing substantially.2
To this end, we’re pleased to share that Open Philanthropy has granted $3M to the NAO over one year to fund a significant scale-up of our wastewater sequencing, targeting three NovaSeq X 25B runs weekly. We’re planning to deploy these funds both in-house and at MU:
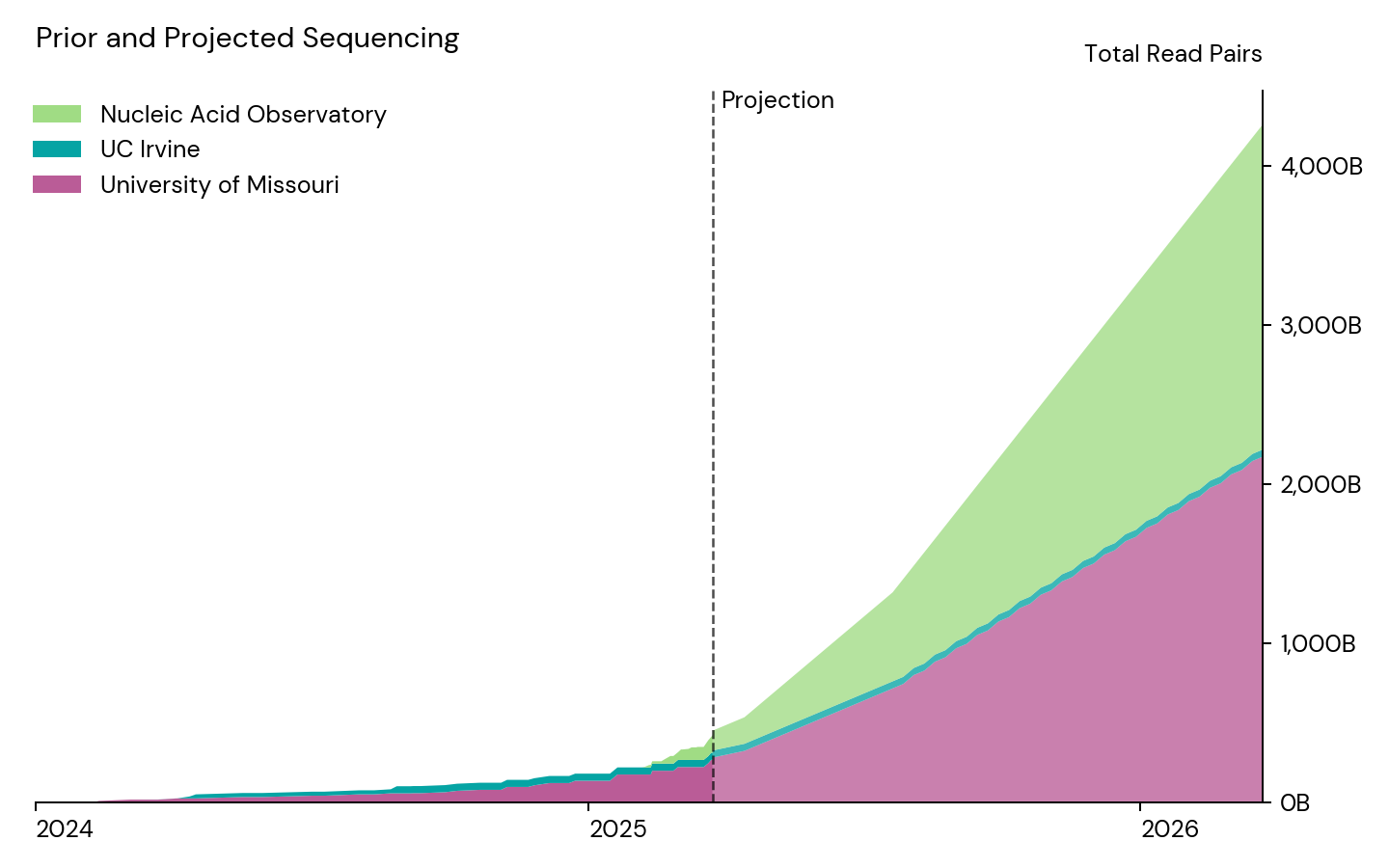
We expect this data to be valuable for a wide range of purposes, including:
Using our existing methods to look for engineered viruses that might be spreading now. While we think such a virus is unlikely to be at large now or over the next year, the consequences of one spreading undetected could be very serious.
Developing additional methods, both reference-based and reference-free, to identify novel pathogens.
Providing a more general supplement to traditional PCR-based public health wastewater surveillance, dramatically expanding the number of pathogens that can be tracked.
Allowing researchers studying the sewer microbiome to better characterize its enormous complexity.
We are grateful for Open Philanthropy’s support, and also to the hard work by our collaborators at MU. Additionally, this project wouldn’t be possible without our partners around the country, in academic labs and treatment plants, who are providing wastewater samples. We’re very excited about what this increased scale will allow us to accomplish over the next year, and as always, we’re excited to collaborate with others who are thinking along similar lines.
Footnotes
Even with centrifugation and filtration to enrich for viral particles, you still have to look through a lot of gut and sewer bacteria to find any. While it’s possible to use targeted methods, such as hybrid capture probes that pull out sequences that match specific known pathogens, a system based on this approach would fail to detect pathogens that didn’t match the probes. Instead, the NAO has worked to optimize sequence-agnostic enrichment methods, and to handle the poor signal-to-noise ratio with sufficiently deep sequencing.↩︎
For example, we estimate that to flag a pathogen that sheds like Influenza A before about 2% of people in the monitored sewersheds have been infected, in the typical case, you’d need three NovaSeq X 25B runs each week.↩︎